Identifying the mechanism that controls Chinese cabbage flowering
- Like
- Digg
- Del
- Tumblr
- VKontakte
- Buffer
- Love This
- Odnoklassniki
- Meneame
- Blogger
- Amazon
- Yahoo Mail
- Gmail
- AOL
- Newsvine
- HackerNews
- Evernote
- MySpace
- Mail.ru
- Viadeo
- Line
- Comments
- Yummly
- SMS
- Viber
- Telegram
- Subscribe
- Skype
- Facebook Messenger
- Kakao
- LiveJournal
- Yammer
- Edgar
- Fintel
- Mix
- Instapaper
- Copy Link
Posted: 23 August 2019 | Bethan Grylls (New Food), Rachael Harper (New Food Magazine) | No comments yet
A new study suggests that Chinese cabbage has its own independent mechanism for flowering.


A team of scientists say they have successfully identified the long noncoding ribonucleic acids (lncRNAs) that are expressed when Chinese cabbage is temporarily exposed to cold temperatures for a period of four weeks.
LncRNAs
LncRNAs are expressed in most plants, with tens of thousands known to be expressed in the model plant Arabidopsis thaliana. Interestingly, lncRNAs that are known to be involved in responding to cold in A. thaliana do not exist in Chinese cabbage. This suggests that this species of cabbage has its own independent mechanism for flowering.
LncRNAs have been linked to vernalisation (when plants need to be exposed to a period of cold treatment in order to flower), which can be seen in many subspecies of Brassica rapa L. including the Chinese cabbage which grows during winter to bloom in spring.
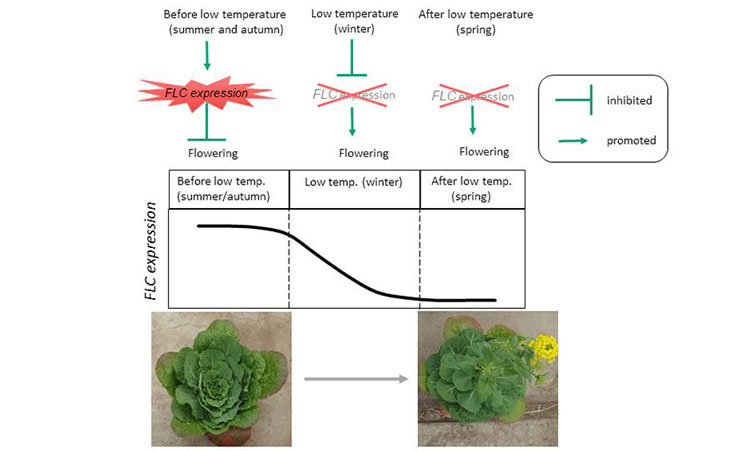

This is floral induction (flowering) in Chinese cabbage (credit: Kobe University).
The research team, led by Namiko Nishida (Kobe University Graduate School of Agricultural Science) and Daniel Shea (Niigata University Graduate School of Science and Technology), extracted total RNAs from Chinese cabbage leaves cultivated for two weeks at 22°C, then Chinese cabbage leaves cultivated at room temperature for two weeks, and finally at a low temperature (4°C) for a further four weeks.
When the team investigated the mRNAs (messenger RNAs) expression patterns induced by low temperatures whose transcription domains overlapped with incRNAs (intronic RNAs, a type of IncRNA) and NATs (natural antisense transcripts, another type of IncRNA), they found similar expression patterns. But, a strong association to vernalisation was not discovered.
What the scientists did uncover were combinations of mRNA and NATs that showed the same altered expression under low temperatures, suggesting that NATs are connected to changes in mRNA expression under cold treatment.
The team searched for the existence of IncRNAs whose expression is induced by cold treatment, focusing on the BrFLC loci. One NAT called BrFLC2as showed the similar structure and expression patterns to COOLAIR (a key IncRNA of A. thaliana), but these similarities were not found in BrFLC1, BrFLC3 or BrFLC5 . In all four BrFLC types, the IncRNAs COLDAIR and COOLWRAP known to recruit the PRC2 complex in A. thaliana was not identified in Chinese cabbage.
This suggests that for Chinese cabbage exposed to low temperatures, the PRC2 complex may be recruited to the FLC locus based on a different mechanism from A. thaliana.
These findings also show that IncRNA expression is not induced in Chinese cabbage from the FLC locus, meaning that it likely follows a different flowering mechanism from the model in A. thaliana.
According the researchers, the next step is to reveal how the PRC2 complex is recruited to the FLC locus under cold treatment. The team believe that identifying the full vernalisation mechanism via FLC genes could contribute to selective breeding of Chinese cabbage.
The findings were published in Scientific Reports.
Related topics
Environment, Processing, Refrigeration and freezing, Research & development, Temperature control









