Loop-mediated isothermal nucleic acid amplification (LAMP) for food microbiology testing
- Like
- Digg
- Del
- Tumblr
- VKontakte
- Buffer
- Love This
- Odnoklassniki
- Meneame
- Blogger
- Amazon
- Yahoo Mail
- Gmail
- AOL
- Newsvine
- HackerNews
- Evernote
- MySpace
- Mail.ru
- Viadeo
- Line
- Comments
- Yummly
- SMS
- Viber
- Telegram
- Subscribe
- Skype
- Facebook Messenger
- Kakao
- LiveJournal
- Yammer
- Edgar
- Fintel
- Mix
- Instapaper
- Copy Link
Posted: 2 April 2017 | Nigel Cook - Fera, Rosario Romero - Fera | No comments yet
Contamination of food products with pathogenic microorganisms is a major concern for food safety. The issue is exacerbated by modern production systems where high volumes, high speed and global distribution are common features. In order to protect consumers, regulations and procedures are in place to control microbiological hazards in the food chain, and testing food samples for a range of bacterial, fungal and viral pathogens is an essential element.

The gold standards in microbiological testing are the methods published by the International Standards Organisation (ISO) (http://www.iso.org/iso/home/about.htm ). The drawback with the ISO methods is that they tend to rely on culture and can therefore be laborious and time-consuming, although they have the advantage of detecting only cells that are viable and able to multiply. However, for bacteria that might be in the hypothesised viable but non-culturable state1 and for those that are harmful even if dead at the time of analysis (e.g. by having released toxins into the food when viable), culture-based methods will not be appropriate. Molecular methods targeting nucleic acid sequences overcome some of these drawbacks and offer speed and higher specificity than culture-based methods.
An extensive range of polymerase chain reaction (PCR)-based nucleic acid detection methods has been developed for the specific detection of numerous foodborne pathogens. A more recent development is the loop-mediated isothermal amplification (LAMP) technique2 and this is emerging as the basis for rapid testing and in-field applications. LAMP is a rapid DNA amplification method that relies on a DNA polymerase that is able to amplify DNA at constant temperature (typically 60oC-65oC). A minimum set of four primers (short DNA strands that specifically determine which DNA fragment will be amplified) is required which recognise six independent regions in the target DNA sequence, achieving high specificity. The amplification products are a variety of DNA molecules with a characteristic loop structure containing copies of the target sequence. An additional pair of primers (loop primers) complementary to the DNA sequence in the loop structure can be included to make the reaction faster.3
Advantages of LAMP over PCR-based techniques
As LAMP amplifies DNA with high efficiency under isothermal conditions, it does not require expensive thermocycler instruments. Assay times are shorter than for PCR, typically 15-60 minutes, depending on the specific polymerase and assay conditions used.
LAMP is highly specific, avoiding both background noise often detected in other nucleic acid amplification techniques and interference by non-target DNA that might be present in the sample.
LAMP is less sensitive to inhibitory substances than PCR, and this can reduce the need to carry out expensive and time-consuming DNA purification procedures prior to testing. This feature is particularly useful for rapid food testing since foods are by nature complex and variable, and so prone to interfere with PCR assays.
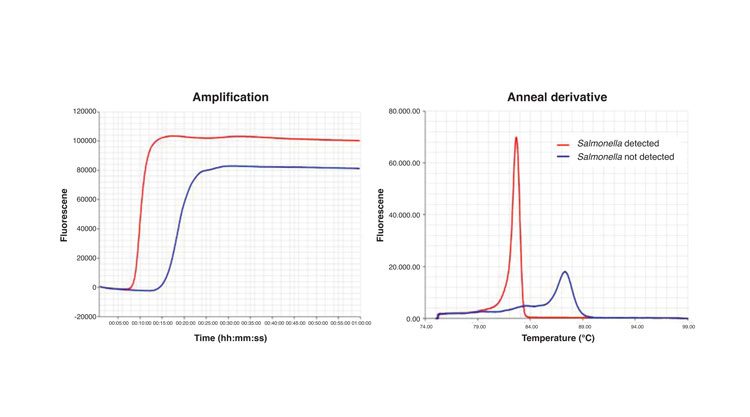

Figure 1: Results of a S. Typhimurium LAMP assay using fluorescence readout. The amplification graph shows the incorporation of the fluorescent dye into the reaction products. The anneal derivative graph shows the characteristic curves of the target and internal amplification control (IAC) amplification products. Red line: results obtained with a sample contaminated with S. Typhimurium; Blue line: results obtained with a sample not containing S. Typhimurium
The assay itself is simple and easy to perform, and does not per se require specialist instruments or a laboratory set up. A simple water bath or heating block is sufficient, which is an advantage for deployment of the technology in remote areas or in low income regions. Detection of amplification products can be by the naked eye or via photometry for turbidity caused by the accumulation of magnesium pyrophosphate precipitate as a by-product of the reaction.4 Alternatively, the reaction can be followed in real-time by measuring fluorescence using dyes which interact with DNA or fluorescent probes (Figure 1), and relatively inexpensive portable instruments are now available for this.
Appropriate controls for LAMP-based methods
In order to avoid any false positive results an internal amplification control (IAC) can be incorporated into the reaction. This control may be a short sequence of DNA with recognition sequences for the same primers as the target but with a different nucleotide sequence elsewhere.5,6 Target DNA and IAC have different physico-chemical properties and therefore, following amplification, they can be easily distinguished from one another by monitoring their behaviour via fluorescence reading after a quick heating step. The amount of IAC for a given assay is optimised to minimise competition with the target DNA. If the sample contains no target DNA, the IAC will be amplified demonstrating that the assay ‘worked’ and no target was present. This control is essential in samples that may contain inhibitors of DNA polymerases. Although LAMP polymerases are very robust, when using crude samples of variable composition, e.g. poultry litter, a certain level of inhibition might occur. The presence of the IAC-specific signal will then serve as confirmation of the negative status of the sample, rather than a failed test which without an IAC would be indistinguishable from a negative result.
Future prospects of LAMP in food microbiology testing
LAMP technology offers great potential for food microbiology and indeed for biological diagnostics in general due to its sensitivity, specificity, speed and
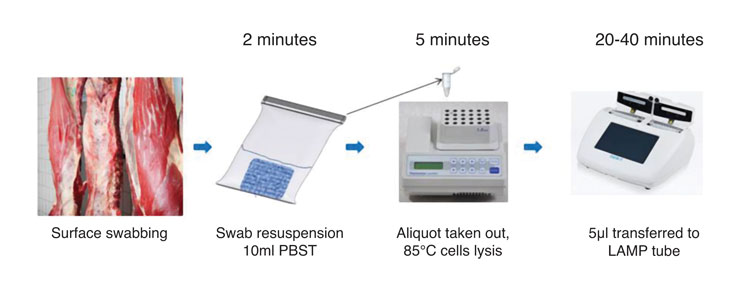

Figure 2: Workflow of a carcass swab testing procedure using LAMP. Results are obtained within one hour
practical applicability (Figure 2 illustrates a rapid procedure for carcass swab testing using LAMP). The technology is clearly emerging as a valuable tool for applications where a rapid answer is required regarding the presence/absence of a microorganism to inform decision making and possible further analysis.
Most of the applications described thus far have the limitation of not being able to discriminate living from dead, or viable but non-culturable cells. Nonetheless, recent reports have described developments in the use of chemicals such as propidium monoazide to block the amplification of DNA from dead cells prior to LAMP.9,10 An alternative option is the detection of mRNA as target since this is only found in viable cells.11
Many other improvements to the LAMP technologies are progressing concerning multiplexing, integration with sample preparation, equipment-free procedures, etc.12 However, an important barrier to acceptance and implementation of molecular-based methods for routine food safety testing is lack of validation. A novel method should be shown to be as effective as the corresponding standard method in order to be fully acceptable as an alternative to the standard, but this has only been done for very few non-proprietorial molecular methods for foodborne pathogen detection. Only one LAMP-based method, that for detection of Salmonella spp. in soya meal, has been validated against the corresponding standard ISO 6579.13 However, the high utility of LAMP warrants consideration of investment to develop efficient modern methods for rapid and sensitive foodborne pathogen detection.
Biographies
Rosario Romero has a long track record in cell biology and protein biochemistry in biomedical research. Since joining Fera in 2011, Rosario has been leading work on applications of protein tools and technologies to food research, with a particular interest in developing proteomics methods for food safety.
Nigel Cook is a graduate of the University of Dundee. At Fera, he studies the transmission of pathogens, particularly enteric viruses, through foods and the environment. He is Councillor of the International Association for Food and Environmental Virology. He was project leader within the standardisation working group ISO TC34 SC9 WG6, developing a standard for detection of Cryptosporidium and Giardia on berry fruits and leafy green vegetables.
References
- Junyan Liu, Rong Zhou, Lin Li, Brian M. Peters, Bing Li, Chii-wann Lin, Tsung-Liang Chuang, Dingqiang Chen, Xihong Zhao, Zhiyong Xiong, Zhenbo Xu, and Mark E. Shirtliff, ‘Viable but Non-Culturable State and Toxin Gene Expression of Enterohemorrhagic Escherichia Coli O157 under Cryopreservation’, Research in Microbiology
- Notomi, H. Okayama, H. Masubuchi, T. Yonekawa, K. Watanabe, N. Amino, and T. Hase, ‘Loop-Mediated Isothermal Amplification of DNA’, Nucleic Acids Res, 28 (2000), E63
- Nagamine, T. Hase, and T. Notomi, ‘Accelerated Reaction by Loop-Mediated Isothermal Amplification Using Loop Primers’, Molecular and Cellular Probes, 16 (2002), 223-29
- Yasuyoshi Mori, Kentaro Nagamine, Norihiro Tomita, and Tsugunori Notomi, ‘Detection of Loop-Mediated Isothermal Amplification Reaction by Turbidity Derived from Magnesium Pyrophosphate Formation’, Biochemical and Biophysical Research Communications, 289 (2001), 150-54
- Martin D’Agostino, Marta Diez-Valcarce, Susana Robles, Begoña Losilla-Garcia, and Nigel Cook, ‘A Loop-Mediated Isothermal Amplification-Based Method for Analysing Animal Feed for the Presence of Salmonella’, Food Analytical Methods, 8 (2015), 2409-16
- Rodriguez-Lazaro, M. D’Agostino, M. Pla, and N. Cook, ‘Construction Strategy for an Internal Amplification Control for Real-Time Diagnostic Assays Using Nucleic Acid Sequence-Based Amplification: Development and Clinical Application’, J Clin Microbiol, 42 (2004), 5832-6
- R. Romero, M. D’Agostino, A. P. Arias, S. Robles, C. F. Casado, L. O. Iturbe, O. G. Lerma, M. Andreou, and N. Cook, ‘An Immunomagnetic Separation/Loop-Mediated Isothermal Amplification Method for Rapid Direct Detection of Thermotolerant Campylobacter Spp. During Poultry Production’, J Appl Microbiol, 120 (2016), 469-77
- Li, Z. Hu, and C. A. Elkins, ‘Detection of Live Escherichia Coli O157:H7 Cells by Pma-Qpcr’, J Vis Exp (2014), e50967
- Chen, F. Wang, J. C. Beaulieu, R. E. Stein, and B. Ge, ‘Rapid Detection of Viable Salmonellae in Produce by Coupling Propidium Monoazide with Loop-Mediated Isothermal Amplification’, Appl Environ Microbiol, 77 (2011), 4008-16
- Y. Youn, O. M. Jeong, B. K. Choi, S. C. Jung, and M. S. Kang, ‘Application of Loop-Mediated Isothermal Amplification with Propidium Monoazide Treatment to Detect Live Salmonella in Chicken Carcasses’, Poult Sci (2016)
- Li, Q. Ying, X. Su, and T. Li, ‘Development and Application of Reverse Transcription Loop-Mediated Isothermal Amplification for Detecting Live Shewanella Putrefaciens in Preserved Fish Sample’, J Food Sci, 77 (2012), M226-30
- Ludwig Niessen, Jie Luo, Carla Denschlag, and Rudi F. Vogel, ‘The Application of Loop-Mediated Isothermal Amplification (Lamp) in Food Testing for Bacterial Pathogens and Fungal Contaminants’, Food Microbiology, 36 (2013), 191-206
- Martin D’Agostino, Susana Robles, Flemming Hansen, Vasileios Ntafis, John Ikonomopoulos, Petros Kokkinos, Avelino Alvarez-Ordonez, Kieran Jordan, Elisabetta Delibato, Elżbieta Kukier, Zbigniew Sieradzki, Krzysztof Kwiatek, Dubravka Milanov, Tamaš Petrović, Patricia Gonzalez-Garcia, David Rodriguez Lazaro, Emily E. Jackson, Stephen J. Forsythe, Lorna O’Brien, and Nigel Cook, ‘Validation of a Loop-Mediated Amplification/ISO 6579-Based Method for Analysing Soya Meal for the Presence of Salmonella Enterica’, Food Analytical Methods, 9 (2016), 2979-85.
Issue
Related topics
Food Safety, Lab techniques, PCR Technology, PCR-based assays









