Hi4GS genomic framework improves wheat yield prediction by 82 percent, helping breeders accelerate development of higher-yielding crop varieties worldwide.

Researchers at Ludong University have developed a new AI-driven genomic framework that improves wheat yield prediction accuracy by more than 82 percent, potentially accelerating the development of higher-yielding crops to support global food security.
Published in The Crop Journal, the Hi4GS (Hybrid Feature Selection for Genomic Selection) framework tackles one of the biggest challenges in modern crop breeding: analysing huge genomic datasets without sacrificing accuracy or increasing computational costs.
Genomic selection has become a core tool in wheat breeding, allowing researchers to predict breeding values using genome-wide markers. However, breeders often struggle with the “small n large p” problem, where datasets contain hundreds of thousands of SNPs (single nucleotide polymorphisms) but relatively few breeding samples. This imbalance can lead to overfitting and inefficient modelling.
The Hi4GS framework streamlines high-dimensional genomic data by isolating the most valuable genetic markers while identifying the biological drivers linked to wheat yield.
Our goal was to move beyond the ‘black box’ nature of traditional genomic models.
By filtering through the noise of tens of thousands of SNPs, Hi4GS allows us to achieve much higher predictive precision with a fraction of the data, while simultaneously uncovering the actual genes that influence yield.”
Shanghui Zhang, the study’s first author
Smarter genomic selection
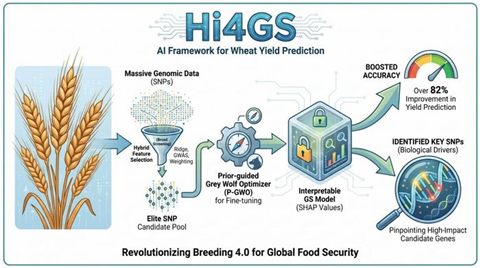
The system combines multiple ranking algorithms with a Prior-guided Grey Wolf Optimizer (P-GWO), which focuses analysis on a pre-screened pool of elite SNP candidates instead of searching entire datasets randomly.
This acts like a navigation map.
By using prior knowledge to guide the starting point, we find the optimal SNP combination faster and more accurately.”
Professor Fa Cui, corresponding author
The researchers also applied SHAP (SHapley Additive exPlanations) values for the first time in genomic wheat prediction, allowing them to measure whether specific SNPs positively or negatively affect yield and reveal interactions between genetic markers.
Testing across 11 yield traits in four wheat datasets showed that genomic selection models using Hi4GS-selected SNPs consistently outperformed models using full SNP datasets. The researchers also found that the SNPs identified by Hi4GS appeared in gene regions at a rate of 9.17 percent, compared with a genomic background rate of 5.61 percent, reinforcing the framework’s ability to isolate functional biological information.
The team believes Hi4GS could support “Breeding 4.0” strategies by helping breeders narrow thousands of markers down to a few dozen high-impact candidate genes, reducing breeding costs and improving efficiency.
Cui said the framework could also be applied beyond wheat to other major crops and animals, supporting genomic-assisted breeding and accelerating the development of higher-yielding varieties worldwide.
The team has released the Hi4GS R package as open-source software on GitHub, making the framework available to agricultural researchers worldwide.












![EcoTrophelia_009[1]](https://d1qyip7ii4bpjm.cloudfront.net/Pictures/380x253/2/2/4/14224_EcoTrophelia_0091.jpg)






No comments yet